PHYLOGEOGRAPHY
Ongoing research efforts by members of our group aim to describe and understand the phylogeographic history of fish in Ireland and elsewhere:
Ongoing research efforts by members of our group aim to describe and understand the phylogeographic history of fish in Ireland and elsewhere:
Phylogeography of Atlantic salmon (Salmo salar) in Ireland as revealed by SNPs
The data set generated from the screening of 388 SNP loci from samples throughout the species range was interrogated and those loci which showed polymorphism in Irish samples (approximately 200 loci) were used to screen approximately 2000 samples from 45 populations sampled in 35 rivers, located in the Republic of Ireland. In addition, samples from rivers in Northern Ireland were obtained and screened for the same loci. Recently, we have supplemented the 35 rivers above with three more rivers from N. Ireland (Bush, Glendun, Shimna) and the Owenmore and Burrishoole Rivers in the west of Ireland. The three additional N. Ireland samples were added in order to resolve the extent of the genetic structure, particularly in the Northeast part of the island. This is the first time that samples taken from across the whole of Ireland have been included in a single dedicated study. The project has involved two screening exercises and the final data have only been made available recently.
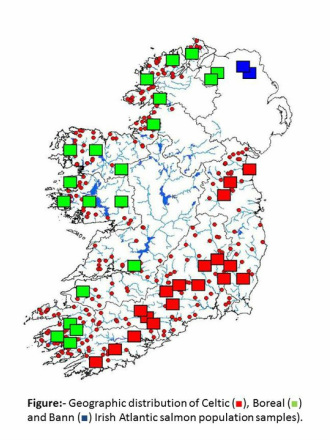
Preliminary results of hierarchical structuring indicate that Irish salmon comprise at least three major phylogenetic lineages which include populations from eastern/southern and western/northern locations, with samples from the river Bann in Northern Ireland representing a further group. The eastern/southern and western/northern groupings appear to correspond to the distributions of the Celtic and Boreal races previously suggested for Ireland. The uniqueness of the Bann samples as revealed by nSNP loci is also supported by microsatellite DNA results and therefore seems unlikely to be a sampling effect. Full hierarchical analysis for further sub-structuring of phylogeographic patterns is ongoing.
The data set generated from the screening of 388 SNP loci from samples throughout the species range was interrogated and those loci which showed polymorphism in Irish samples (approximately 200 loci) were used to screen approximately 2000 samples from 45 populations sampled in 35 rivers, located in the Republic of Ireland. In addition, samples from rivers in Northern Ireland were obtained and screened for the same loci. Recently, we have supplemented the 35 rivers above with three more rivers from N. Ireland (Bush, Glendun, Shimna) and the Owenmore and Burrishoole Rivers in the west of Ireland. The three additional N. Ireland samples were added in order to resolve the extent of the genetic structure, particularly in the Northeast part of the island. This is the first time that samples taken from across the whole of Ireland have been included in a single dedicated study. The project has involved two screening exercises and the final data have only been made available recently.
Preliminary results of hierarchical structuring indicate that Irish salmon comprise at least three major phylogenetic lineages which include populations from eastern/southern and western/northern locations, with samples from the river Bann in Northern Ireland representing a further group. The eastern/southern and western/northern groupings appear to correspond to the distributions of the Celtic and Boreal races previously suggested for Ireland. The uniqueness of the Bann samples as revealed by nSNP loci is also supported by microsatellite DNA results and therefore seems unlikely to be a sampling effect. Full hierarchical analysis for further sub-structuring of phylogeographic patterns is ongoing.

Phylogeographic structure of brown trout (Salmo trutta) in Britain and Ireland: glacial refugia, post-glacial colonisation, and origins of sympatric populations
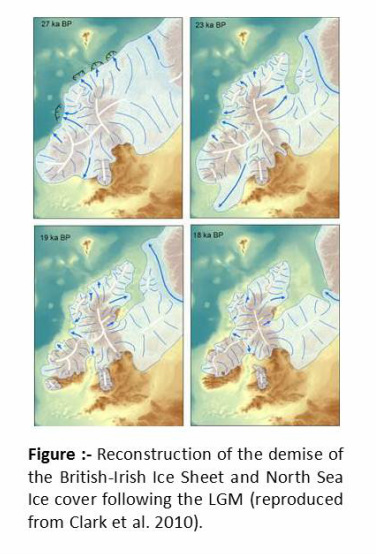
The brown trout displays extensive genetic, ecological, morphological and life-history variation, which has resulted in a long-standing debate on the evolutionary origins and taxonomic implications of this variability. During the Last Glacial Maxima (LGM), which reached a maximum some 23000 to 18000 years before the present, ice covered much of Britain and Ireland, although some peripheral regions such as the south and west of Ireland, and south of England remained free from glaciations. In addition, as a result of the lowered sea level, the land mass was considerably extended in comparison with today providing possible refugia in areas that are currently submerged.
Brown trout are potentially anadromous and this would have allowed both rapid range retreats into refugia during glacial periods, and subsequent expansion during interglacial periods. Similarly, the sea would not have presented a barrier to postglacial colonisation of current freshwater catchments. Thus most, if not all, native brown trout populations in Britain and Ireland, irrespective of current life history, colonised as anadromous (sea) trout.
Anadromy has allowed continued gene flow between populations in adjacent catchments. These interlinked factors suggest that the phylogeographic structure of rown trout in NW Europe is likely to be more complex than that described for many other organisms. Overall, there have been repeated opportunities for genetic divergence in allopatric refuges, followed by interbreeding or by varying degrees of reproductive isolation, on secondary contact after postglacial colonisation. While the consensus is that postglacial colonisation involved multiple lineages originating from separate refuges, hypotheses differ in terms of the number, origin and dispersal dynamics of these lineages. More extensive sampling and increased phylogeographic resolution are therefore necessary to clarify the evolutionary history of brown trout in this region, especially given that, to date, only limited studies have been undertaken in Britain and Ireland
In several waters throughout the range of the species, two or more populations of brown trout have been found to co-occur. One of the best studied examples is the sympatric gillaroo, sonaghen and ferox populations in Lough Melvin, NW Ireland (reviewed by Ferguson, 2004). Based on their genetic, morphological and ecological differentiation, Ferguson (2004) proposed reinstatement of the 19th Century scientific names of gillaroo Salmo stomachicus Günther 1866, sonaghen Salmo nigripinnis Günther 1866, and ferox Salmo ferox Jardine 1935. However, a fuller understanding of the evolutionary origins and homologies to other brown trout populations is necessary for validation of this classification, and for implementation of conservation plans. In particular, it is essential to elucidate whether these sympatric populations in Melvin are the result of colonisation by two or more genetically distinct lineages, or of colonisation by a single lineage followed by splitting within the lake, under the influence of selective forces.
We have carried out a comprehensive collection of brown trout specimens (N = 3636) from 81 sites throughout Britain and Ireland. All samples were screened using PCR-RFLP analysis of four mitochondrial DNA segments covering over 10000 base-pairs (ca 60%) of the complete brown trout mitochondrial genome. We indentified a total of 26 mtDNA haplotypes in this region, considerably extending the phylogenetic resolution relative to previous studies. The haplotypes could be nested into seven 2-step clades. Although there was clear geographical structure to the occurrence of derived clades, admixture among ancestral clades was extensive throughout the studied area. Based on data presented here and published data, postglacial colonisation of Britain and Ireland most likely involved brown trout from at least five potential glacial refuges. Probable locations for such refugia were: south of England / western France; east of the Baltic Sea; western Ireland; Celtic Sea; and North Sea. A relevant feature of the data was that some populations contained mixtures of highly divergent clades. This 'type II' phylogeographic pattern is uncommon in nature. Clade intermixing is likely to have taken place during earlier inter-glacials as well as since the Last Glacial Maximum. The anadromous life history of many trout populations has probably also contributed to clade mixing.
The brown trout displays extensive genetic, ecological, morphological and life-history variation, which has resulted in a long-standing debate on the evolutionary origins and taxonomic implications of this variability. During the Last Glacial Maxima (LGM), which reached a maximum some 23000 to 18000 years before the present, ice covered much of Britain and Ireland, although some peripheral regions such as the south and west of Ireland, and south of England remained free from glaciations. In addition, as a result of the lowered sea level, the land mass was considerably extended in comparison with today providing possible refugia in areas that are currently submerged.
Brown trout are potentially anadromous and this would have allowed both rapid range retreats into refugia during glacial periods, and subsequent expansion during interglacial periods. Similarly, the sea would not have presented a barrier to postglacial colonisation of current freshwater catchments. Thus most, if not all, native brown trout populations in Britain and Ireland, irrespective of current life history, colonised as anadromous (sea) trout.
Anadromy has allowed continued gene flow between populations in adjacent catchments. These interlinked factors suggest that the phylogeographic structure of rown trout in NW Europe is likely to be more complex than that described for many other organisms. Overall, there have been repeated opportunities for genetic divergence in allopatric refuges, followed by interbreeding or by varying degrees of reproductive isolation, on secondary contact after postglacial colonisation. While the consensus is that postglacial colonisation involved multiple lineages originating from separate refuges, hypotheses differ in terms of the number, origin and dispersal dynamics of these lineages. More extensive sampling and increased phylogeographic resolution are therefore necessary to clarify the evolutionary history of brown trout in this region, especially given that, to date, only limited studies have been undertaken in Britain and Ireland
In several waters throughout the range of the species, two or more populations of brown trout have been found to co-occur. One of the best studied examples is the sympatric gillaroo, sonaghen and ferox populations in Lough Melvin, NW Ireland (reviewed by Ferguson, 2004). Based on their genetic, morphological and ecological differentiation, Ferguson (2004) proposed reinstatement of the 19th Century scientific names of gillaroo Salmo stomachicus Günther 1866, sonaghen Salmo nigripinnis Günther 1866, and ferox Salmo ferox Jardine 1935. However, a fuller understanding of the evolutionary origins and homologies to other brown trout populations is necessary for validation of this classification, and for implementation of conservation plans. In particular, it is essential to elucidate whether these sympatric populations in Melvin are the result of colonisation by two or more genetically distinct lineages, or of colonisation by a single lineage followed by splitting within the lake, under the influence of selective forces.
We have carried out a comprehensive collection of brown trout specimens (N = 3636) from 81 sites throughout Britain and Ireland. All samples were screened using PCR-RFLP analysis of four mitochondrial DNA segments covering over 10000 base-pairs (ca 60%) of the complete brown trout mitochondrial genome. We indentified a total of 26 mtDNA haplotypes in this region, considerably extending the phylogenetic resolution relative to previous studies. The haplotypes could be nested into seven 2-step clades. Although there was clear geographical structure to the occurrence of derived clades, admixture among ancestral clades was extensive throughout the studied area. Based on data presented here and published data, postglacial colonisation of Britain and Ireland most likely involved brown trout from at least five potential glacial refuges. Probable locations for such refugia were: south of England / western France; east of the Baltic Sea; western Ireland; Celtic Sea; and North Sea. A relevant feature of the data was that some populations contained mixtures of highly divergent clades. This 'type II' phylogeographic pattern is uncommon in nature. Clade intermixing is likely to have taken place during earlier inter-glacials as well as since the Last Glacial Maximum. The anadromous life history of many trout populations has probably also contributed to clade mixing.
Tracking the evolutionary pathways of the Atlantic salmon (Salmo salar) in Ireland: disentangling postglacial colonisation history from contemporary factors
Verspoor et al. (2005) based on the RFLP analyses of the 16S mtDNA, indentified seven major regional clusters among Eastern Atlantic salmon populations. In that study, population samples from Britain and Ireland were clustered as a single group. This result is in contrast with those of previous investigations (Payne et al. 1971, Child et al.1976) that reported a latitudinal cline in allelic frequencies at the nuclear transferrin locus, which appeared to identify two races of salmon in this region. The populations from rivers draining into the Celtic and the Irish Seas were referred to as the “Celtic” race while more northerly populations were assigned to a putative “Boreal” race. The authors suggested that these races would have resulted from isolation in different glacial refugia.
More recently, Griffiths et al. (2010), based on nuclear microsatellite marker loci, indicated that Atlantic salmon in Britain (in particular) and Ireland are subdivided into a number of regional genetic clusters. For the Irish samples included in that study, there appeared, despite limited sampling coverage, to be some separation between southern and northern-western populations. Preliminary results from the ongoing EU-sponsored SALSEA-MERGE project, which includes more substantial sample coverage throughout the British Isles, seems to confirm the existence of genetically distinct population clusters in this region. Notwithstanding the scientific relevance in terms of understanding evolutionary processes, from both conservation and management perspectives, there is a clear need for a study examining the relative importance of colonisation history and contemporary factors, determining the genetic features and patterns of population structure observed in Irish salmon.
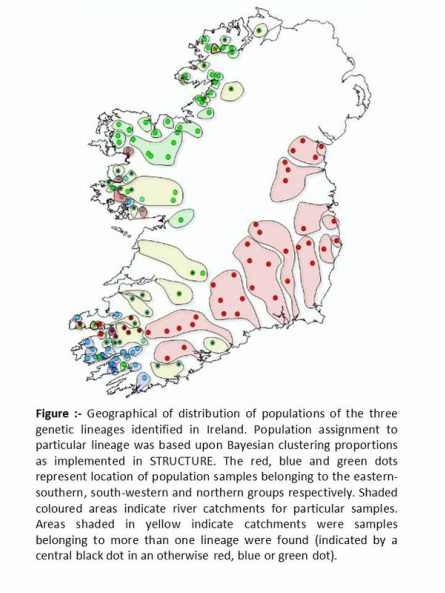
The major deficiency of previous studies was related to sample coverage (relatively few populations sampled from the area of interest) and in some cases limited genetic data. The comprehensive genetic data set generated for the National Genetic Stock Identification baseline, which includes genotypic microsatellite data across 15 loci for over 8000 specimens representing 136 population samples in 80 salmon rivers from throughout the Republic of Ireland circumvents these problems. Thus, it provides a unique opportunity to address the question. Using a combination of Bayesian and non-parametric clustering algorithms, in addition to standard statistical genetics analyses, we have discovered that Irish salmon populations are divided not into two lineages, but into three unambiguous evolutionary lineages The distribution of populations derived from these lineages shows a remarkable geographic discreteness. The three lineages predominate in eastern-southern, south-western and northern rivers, respectively (see adjacent figure). The eastern-southern lineage appears to correspond to the Payne et al. “Celtic” race, whereas the distribution of the other two lineages seems to be similar to that of their “Boreal” race. Some areas of overlap between the distributions of the lineages may represent zones of secondary contact or introgression.
We are currently combining the data from additional analysis involving estimating levels of migration; effective population sizes and degree of population differentiation with that of the perceived colonisation history, in order to disentangle the evolutionary processes that shape populations. We are also investigating factors promoting the maintenance of these distinct lineages such hybrid zones and/or zones of secondary contact.
Verspoor et al. (2005) based on the RFLP analyses of the 16S mtDNA, indentified seven major regional clusters among Eastern Atlantic salmon populations. In that study, population samples from Britain and Ireland were clustered as a single group. This result is in contrast with those of previous investigations (Payne et al. 1971, Child et al.1976) that reported a latitudinal cline in allelic frequencies at the nuclear transferrin locus, which appeared to identify two races of salmon in this region. The populations from rivers draining into the Celtic and the Irish Seas were referred to as the “Celtic” race while more northerly populations were assigned to a putative “Boreal” race. The authors suggested that these races would have resulted from isolation in different glacial refugia.
More recently, Griffiths et al. (2010), based on nuclear microsatellite marker loci, indicated that Atlantic salmon in Britain (in particular) and Ireland are subdivided into a number of regional genetic clusters. For the Irish samples included in that study, there appeared, despite limited sampling coverage, to be some separation between southern and northern-western populations. Preliminary results from the ongoing EU-sponsored SALSEA-MERGE project, which includes more substantial sample coverage throughout the British Isles, seems to confirm the existence of genetically distinct population clusters in this region. Notwithstanding the scientific relevance in terms of understanding evolutionary processes, from both conservation and management perspectives, there is a clear need for a study examining the relative importance of colonisation history and contemporary factors, determining the genetic features and patterns of population structure observed in Irish salmon.
The major deficiency of previous studies was related to sample coverage (relatively few populations sampled from the area of interest) and in some cases limited genetic data. The comprehensive genetic data set generated for the National Genetic Stock Identification baseline, which includes genotypic microsatellite data across 15 loci for over 8000 specimens representing 136 population samples in 80 salmon rivers from throughout the Republic of Ireland circumvents these problems. Thus, it provides a unique opportunity to address the question. Using a combination of Bayesian and non-parametric clustering algorithms, in addition to standard statistical genetics analyses, we have discovered that Irish salmon populations are divided not into two lineages, but into three unambiguous evolutionary lineages The distribution of populations derived from these lineages shows a remarkable geographic discreteness. The three lineages predominate in eastern-southern, south-western and northern rivers, respectively (see adjacent figure). The eastern-southern lineage appears to correspond to the Payne et al. “Celtic” race, whereas the distribution of the other two lineages seems to be similar to that of their “Boreal” race. Some areas of overlap between the distributions of the lineages may represent zones of secondary contact or introgression.
We are currently combining the data from additional analysis involving estimating levels of migration; effective population sizes and degree of population differentiation with that of the perceived colonisation history, in order to disentangle the evolutionary processes that shape populations. We are also investigating factors promoting the maintenance of these distinct lineages such hybrid zones and/or zones of secondary contact.
The phylogeography of the three-spine stickleback (Gasterosteus aculeatus L.) in Britain and Ireland
Within the last two decades the three-spine stickleback has become an important model species for rapid, parallel evolution. Morphological, ecological and physiological divergence between ancestral marine and derived freshwater stickleback populations has thus evolved rapidly and in parallel. With an increasing understanding of the genetic architecture underlying freshwater adaptation in stickleback, there is a clear need for establishing baseline knowledge of stickleback postglacial recolonisation scenarios and spatial genetic variation.
Mäkinen et al. (2006) and Mäkinen and Merilä (2007) previously reconstructed a broad European phylogeography for three-spine stickleback using microsatellite markers and mitochondrial DNA (mtDNA). They established evidence for four major mitochondrial lineages within Europe, with major European and Trans-Atlantic clades found in the British Isles. Microsatellite analysis was consistent with these lineages but also suggested more subtle structuring; for example within the British Isles, populations were grouped either as being of North Atlantic or North Sea origin.
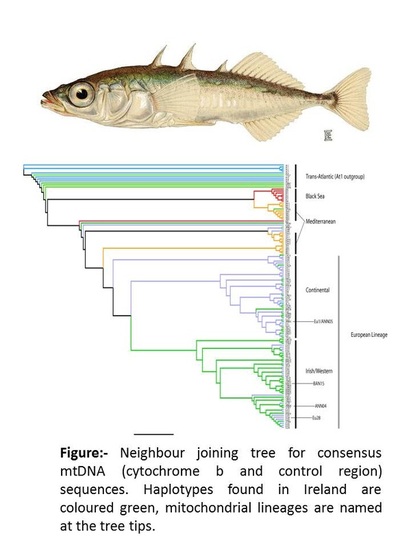
While the findings of these studies remain informative, they were based on only five populations from the UK and none from Ireland. Indeed, to date very limited information is available on three-spine stickleback in Ireland exists. Thus, given the westerly distribution, Holocene isolation and well-understood glacial history of the British Isles, a higher resolution phylogeographical study of stickleback populations in the region is highly desirable. With little knowledge of the distribution of three-spine stickleback in Ireland, an initial baseline survey was conducted and individuals were collected from 31 sample sites. Approximately 10 individuals from each population were then sequenced for two mitochondrial genes (cytochrome B and control region) and a neighbour joining tree was constructed using consensus sequences. Irish populations were analysed alongside European sequences from Mäkinen and Merilä (2007).
These initial data indicated that Ireland represented a zone of secondary contact for both European and Trans-Atlantic mitochondrial lineages. However, a number of unique haplotypes were discovered in Irish populations. These appear to be shallowly diverged from the most frequent European haplotype groups and are positioned alongside populations from Western France and Spain (above figure). Thus it seems that the European mitochondrial lineage suggested by Mäkinen and Merilä (2007) may in fact harbour shallow divergence between Western and Eastern Europe; most likely as a result of recolonisation through the North Atlantic and North Sea basins respectively.
Based on these initial results, coverage of stickleback populations was extended across Ireland and Britain. Thus, in 2010 a more extensive sampling survey was conducted in conjunction with Inland Fisheries Ireland, the Marine Institute and collaborative research groups. Approximately 10 individuals from over 50 study sites have been sequenced for two mitochondrial genes (cytochrome b and control region) and genotyped at nine neutral microsatellite loci. Data have now been compiled to reconstruct a phylogeny of these populations, using both marker sets in a Bayesian approach. Additionally, by drawing on a recently published synopsis on the glacial chronology and extent of the British Irish Ice Sheet we have used a statistical approach through Bayesian model selection to find the most appropriate phylogeographic fit to the data.
Within the last two decades the three-spine stickleback has become an important model species for rapid, parallel evolution. Morphological, ecological and physiological divergence between ancestral marine and derived freshwater stickleback populations has thus evolved rapidly and in parallel. With an increasing understanding of the genetic architecture underlying freshwater adaptation in stickleback, there is a clear need for establishing baseline knowledge of stickleback postglacial recolonisation scenarios and spatial genetic variation.
Mäkinen et al. (2006) and Mäkinen and Merilä (2007) previously reconstructed a broad European phylogeography for three-spine stickleback using microsatellite markers and mitochondrial DNA (mtDNA). They established evidence for four major mitochondrial lineages within Europe, with major European and Trans-Atlantic clades found in the British Isles. Microsatellite analysis was consistent with these lineages but also suggested more subtle structuring; for example within the British Isles, populations were grouped either as being of North Atlantic or North Sea origin.
While the findings of these studies remain informative, they were based on only five populations from the UK and none from Ireland. Indeed, to date very limited information is available on three-spine stickleback in Ireland exists. Thus, given the westerly distribution, Holocene isolation and well-understood glacial history of the British Isles, a higher resolution phylogeographical study of stickleback populations in the region is highly desirable. With little knowledge of the distribution of three-spine stickleback in Ireland, an initial baseline survey was conducted and individuals were collected from 31 sample sites. Approximately 10 individuals from each population were then sequenced for two mitochondrial genes (cytochrome B and control region) and a neighbour joining tree was constructed using consensus sequences. Irish populations were analysed alongside European sequences from Mäkinen and Merilä (2007).
These initial data indicated that Ireland represented a zone of secondary contact for both European and Trans-Atlantic mitochondrial lineages. However, a number of unique haplotypes were discovered in Irish populations. These appear to be shallowly diverged from the most frequent European haplotype groups and are positioned alongside populations from Western France and Spain (above figure). Thus it seems that the European mitochondrial lineage suggested by Mäkinen and Merilä (2007) may in fact harbour shallow divergence between Western and Eastern Europe; most likely as a result of recolonisation through the North Atlantic and North Sea basins respectively.
Based on these initial results, coverage of stickleback populations was extended across Ireland and Britain. Thus, in 2010 a more extensive sampling survey was conducted in conjunction with Inland Fisheries Ireland, the Marine Institute and collaborative research groups. Approximately 10 individuals from over 50 study sites have been sequenced for two mitochondrial genes (cytochrome b and control region) and genotyped at nine neutral microsatellite loci. Data have now been compiled to reconstruct a phylogeny of these populations, using both marker sets in a Bayesian approach. Additionally, by drawing on a recently published synopsis on the glacial chronology and extent of the British Irish Ice Sheet we have used a statistical approach through Bayesian model selection to find the most appropriate phylogeographic fit to the data.
Global population structure, phylogeography and historical demography of the blue shark, Prionace glauca – inferences from mitochondrial DNA analysis
The blue shark is the most widespread and abundant of the extant pelagic sharks. Despite its ecological and economic importance, little is known of this species stock structure on a global scale. Blue sharks possess a remarkable life history, which includes complex migratory patterns involving segregation by size and sex immediately after birth and extensive cyclical migrations between breeding grounds and pupping areas. These movements are intrinsically linked with the major oceanic current systems and also with seasonal shifts in zones of high productivity. It has also been found that the female of many species of shark exhibit philopatry to breeding and/or parturition sites. Furthermore, gene flow between relatively distant locations seems to be predominantly mediated by roaming males. Evidence for these sex specific behaviours (reviewed by Hueter et al., 2005) comes directly from tagging/tracking studies, which demonstrate annual/biannual returns to specific locations, and indirectly from genetic studies that have shown significant discrepancies in the level of genetic divergence as measured from mtDNA (maternally inherited segregation) in comparison to equivalent nuclear data
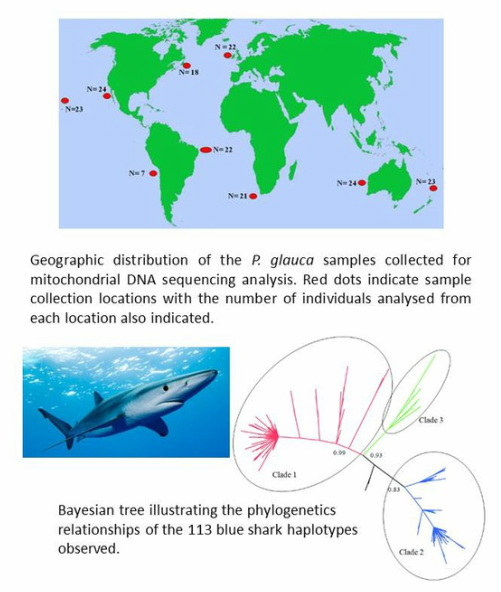
Past paleoclimatic events, which have resulted in oceanographic shifts, may also have had a major impact on the phylogeographic and demographic history of this species and is likely to have had an influence on the population structure we observed today. A sample of 164 blue shark tissue samples were selected nine different locations representing the geographic location of the species (above figure). We used five species-specific mtDNA PCR primer sets, which enabled the amplification of 3,107 base pair from the blue shark mtDNA genome. Resulting products were sequenced and the data obtained subsequently used in population genetics and phylogeographic analyses. In order to improve the precision of times estimates for divergence above, we have recently completed the sequencing of 32 entire mitochondrial DNA genomes representative of each of the three clades identified.
Our phylogeographic analyses support the existence of three well defined maternal clades (above figure). While we found no major geographical patterns with regards to contemporary distribution of the mtDNA haplotypes belonging to each of these clades, we observed significant variation in haplotypic frequency distribution among samples from different ocean basins, the North Atlantic been the most differentiated. It would appear from our complementary population genetic analysis that female blue shark display site fidelity with to breeding grounds. The estimated time of expansion for clades 1 and 2 were approximately 1 million and 773 thousand years respectively, while the estimated time for the whole data set was 1.42 million years.
Policy development/economic implications
Sharks are fish species whose conservation falls within the domain of the Common Fishery Policy (CFP). The Community plan is based on the Council Regulation (EC) N° 2371/2002 of 20 December 2002 for the conservation and sustainable exploitation of fisheries resources under the CFP. In addition, within EC legislation there are provisions dealing with output management, technical measures, control, fleet and market policy which could be effective in ensuring a sustainable exploitation of elasmobranchs. There is major ongoing review of policy and regulations as it pertains to the management of sharks. As one most seriously impacted species from fisheries bycatch, the blue shark is expected to be provided special status within that legislation. Our genetics studies will contribute to these deliberations.
Predicted impact on management and scientific community
From a management perspective, should it be shown that sharks display fidelity to particular geographic locations, as opposed to roaming and dispersing throughout their distribution ranges, the impact of fisheries removals and habitat alterations on particular areas could be profound. The scientific impact of this study is its ability to examine evolutionary events that took place millions of years ago in contrast to the majority of phylogeographic investigations, which look at more recent events associated with the last glaciation a mere 20,000 years ago.
The blue shark is the most widespread and abundant of the extant pelagic sharks. Despite its ecological and economic importance, little is known of this species stock structure on a global scale. Blue sharks possess a remarkable life history, which includes complex migratory patterns involving segregation by size and sex immediately after birth and extensive cyclical migrations between breeding grounds and pupping areas. These movements are intrinsically linked with the major oceanic current systems and also with seasonal shifts in zones of high productivity. It has also been found that the female of many species of shark exhibit philopatry to breeding and/or parturition sites. Furthermore, gene flow between relatively distant locations seems to be predominantly mediated by roaming males. Evidence for these sex specific behaviours (reviewed by Hueter et al., 2005) comes directly from tagging/tracking studies, which demonstrate annual/biannual returns to specific locations, and indirectly from genetic studies that have shown significant discrepancies in the level of genetic divergence as measured from mtDNA (maternally inherited segregation) in comparison to equivalent nuclear data
Past paleoclimatic events, which have resulted in oceanographic shifts, may also have had a major impact on the phylogeographic and demographic history of this species and is likely to have had an influence on the population structure we observed today. A sample of 164 blue shark tissue samples were selected nine different locations representing the geographic location of the species (above figure). We used five species-specific mtDNA PCR primer sets, which enabled the amplification of 3,107 base pair from the blue shark mtDNA genome. Resulting products were sequenced and the data obtained subsequently used in population genetics and phylogeographic analyses. In order to improve the precision of times estimates for divergence above, we have recently completed the sequencing of 32 entire mitochondrial DNA genomes representative of each of the three clades identified.
Our phylogeographic analyses support the existence of three well defined maternal clades (above figure). While we found no major geographical patterns with regards to contemporary distribution of the mtDNA haplotypes belonging to each of these clades, we observed significant variation in haplotypic frequency distribution among samples from different ocean basins, the North Atlantic been the most differentiated. It would appear from our complementary population genetic analysis that female blue shark display site fidelity with to breeding grounds. The estimated time of expansion for clades 1 and 2 were approximately 1 million and 773 thousand years respectively, while the estimated time for the whole data set was 1.42 million years.
Policy development/economic implications
Sharks are fish species whose conservation falls within the domain of the Common Fishery Policy (CFP). The Community plan is based on the Council Regulation (EC) N° 2371/2002 of 20 December 2002 for the conservation and sustainable exploitation of fisheries resources under the CFP. In addition, within EC legislation there are provisions dealing with output management, technical measures, control, fleet and market policy which could be effective in ensuring a sustainable exploitation of elasmobranchs. There is major ongoing review of policy and regulations as it pertains to the management of sharks. As one most seriously impacted species from fisheries bycatch, the blue shark is expected to be provided special status within that legislation. Our genetics studies will contribute to these deliberations.
Predicted impact on management and scientific community
From a management perspective, should it be shown that sharks display fidelity to particular geographic locations, as opposed to roaming and dispersing throughout their distribution ranges, the impact of fisheries removals and habitat alterations on particular areas could be profound. The scientific impact of this study is its ability to examine evolutionary events that took place millions of years ago in contrast to the majority of phylogeographic investigations, which look at more recent events associated with the last glaciation a mere 20,000 years ago.

